1.表观遗传调控
-
Liang et al. Distinct dynamics of parental 5-hydroxymethylcytosine during human preimplantation development regulate early lineage gene expression. Nat Cell Biol (2024)
PMID: 39080410
-
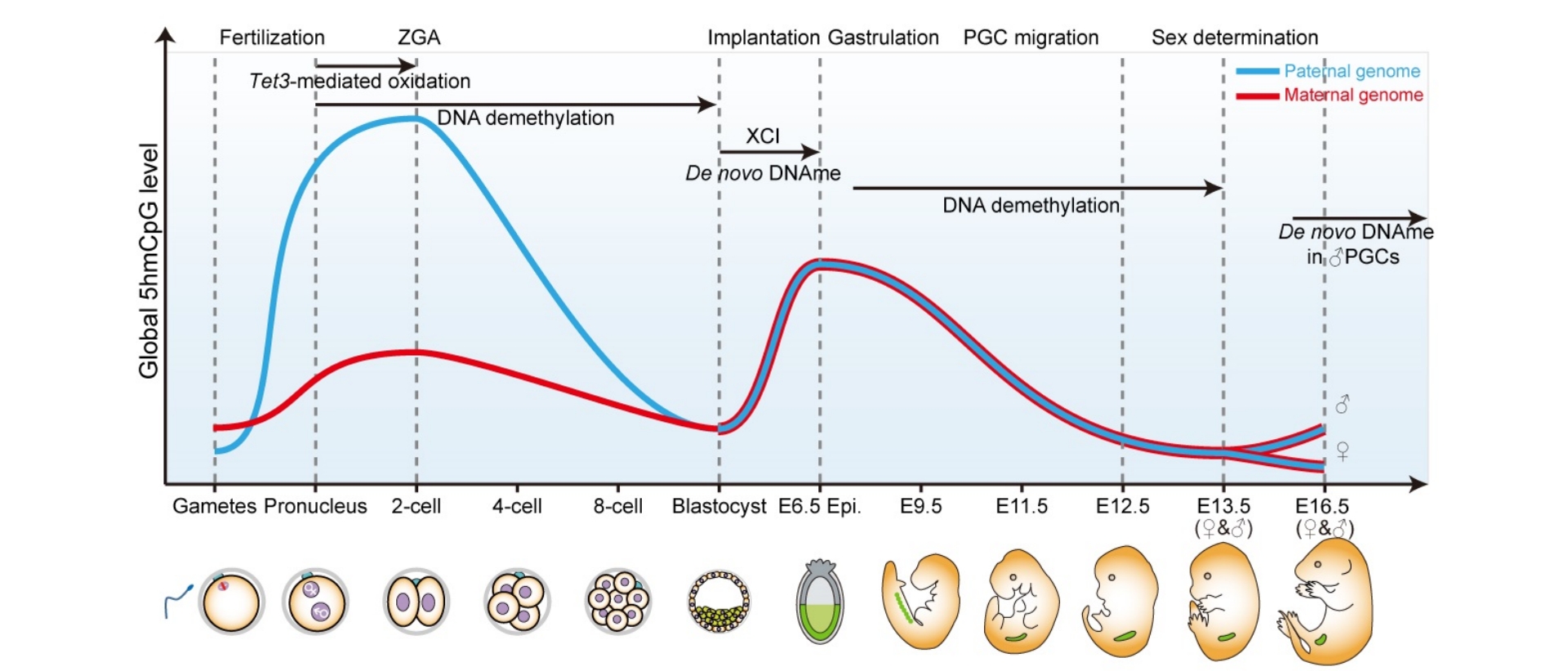
Yan et al. Dynamics of DNA hydroxymethylation and methylation during mouse embryonic and germline development. Nat Genet (2023)
PMID: 36539615
-
Yan et al. Decoding dynamic epigenetic landscapes in human oocytes using single-cell multi-omics sequencing. Cell Stem Cell (2021)
PMID: 33957080
-
Gu et al. Integrative single-cell analysis of transcriptome, DNA methylome and chromatin accessibility in mouse oocytes. Cell Res (2019)
PMID: 30560925
-
Li et al. Single-cell multi-omics sequencing of human early embryos. Nat Cell Biol (2018)
PMID: 29915357
-
Guo et al. Single-cell multi-omics sequencing of mouse early embryos and embryonic stem cells. Cell Res (2017)
PMID: 28621329
-
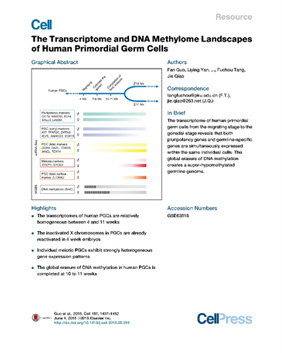
Guo et al. The transcriptome and DNA methylome landscapes of human primordial germ cells. Cell (2015)
PMID: 26046443
-
Guo et al. Active and passive demethylation of male and female pronuclear DNA in the mammalian zygote. Cell Stem Cell (2014)
PMID: 25220291
-
Gu et al. The role of Tet3 DNA dioxygenase in epigenetic reprogramming by oocytes. Nature (2011)
PMID: 21892189









































 联系人:曹羽
联系人:曹羽 电子邮件:caoyu@ioz.ac.cn
电子邮件:caoyu@ioz.ac.cn 通讯地址:北京市朝阳区北辰西路1号院5号
通讯地址:北京市朝阳区北辰西路1号院5号